What is CRISPR? Where did it come from and how does it work?
CRISPR is a modern, high-tech method of editing genes, but it’s based on a simple natural defence system found in many bacteria. Like higher organisms, bacteria also have to cope with a number of viruses and plasmids that can invade and take over their cells. The bacteria maintain a sort of genetic library within their DNA, containing key bits of gene sequences from past encounters with pathogens they might normally have to contend with. CRISPR refers to Clustered Regularly Interspaced Short Palindromic Repeats, which is basically what the bacterial libraries use to catalogue various viral and plasmid threats. These sequences, in conjunction with special enzymes, can serve as templates for precise molecular “scissors” that hunt down and cut DNA or RNA to destroy intracellular invaders before they can replicate. The trick to avoid any self-inflicted damage to the genetic sequence on file in the bacterium’s library is that, in addition to the specific viral genetic sequence, the scissors also look for what is called protospacer adjacent motif (PAM), which is found in the invader’s genetic material but not in the bacterium’s.
Once scientists learned how to programme the molecular scissors of CRISPR using their own custom-designed genetic sequences rather than those from a bacterium’s library, they could target specific points on DNA strands to turn off, or “knock out” certain genes in almost any organism. This type of whack-a-mole research still continues in many aquatic species, especially with regard to health and immunity questions. But, after a brief period of somewhat exploratory research based on a “knock stuff out and see what happens” approach, a number of “maybe we can turn stuff on” applications began to emerge.
If modifications from gene editing are to be heritable, however, they usually must take place very early in the development of an animal or plant (ideally at the one-cell stage) so that they can be incorporated in its gametes once it matures. With this caveat comes the need for specialised methods like microinjection to deliver the CRISPR constructs and enzymes on a microscopic scale. An annoying side-effect of this approach is that the CRISPR effect can continue within the developing embryo well beyond the one-cell stage, often resulting in mosaicism (when various cells display different genetic makeups).
A review of the science and the step-by-step details of how CRISPR has advanced are beyond the scope of this article (and the expertise of this author, for that matter), but suffice it to say that the potential applications for the technology now seem limited only by the imaginations of the scientists that have mastered it. Some examples include altering mosquitos so they cannot find human targets, breeding coffee beans with all the flavour and none of the caffeine, developing hangover-free wine with health benefits, creating tomatoes with the same spice producing genes as chili peppers (the genes are already present, just not expressed), molecular sleuthing for pathogens in natural or man-made environments and even bringing back the extinct wooly mammoth (well, actually creating a sort of mammoth/elephant hybrid, depending how one looks at it).
What's been done so far?
To date, most CRISPR research in farmed aquatic organisms reflects the same emphases as more traditional genetic improvement initiatives, namely growth, disease resistance and sterility, although some interesting work has also been done with regard to colouration patterns in various fishes. Problems with applying the technology to complex traits such as growth and disease resistance remain, since a number of genes are involved and many of them might require editing to attain desired outcomes. Fortunately advances in genomic selection continue in many aquatic species and this may provide shortcuts for the application of CRISPR, but significant work will be required to determine which versions of which genes should be targeted through editing. Editing to insert artificial alleles or genetic sequences from other species is now possible, but it will most likely face the same resistance observed toward the commercialisation of transgenic aquatic organisms.

Her keynote address on the subject at Aquaculture Europe in Dubrovnik in 2017 attracted over 500 delegates. © Rob Fletcher
In salmonids, much of the work to date has focused on Atlantic salmon. While CRISPR can be used to produce sterile salmon by targeting the “dead end” gene, more effort will be required to apply this approach commercially because by default it cannot be transmitted to subsequent generations. Nonetheless, prior work with laboratory species (zebrafish and medaka) suggests that a method to restore fertility for breeding stocks might be possible. In other research, the dead end gene has been used to produce sterile fish which can then be implanted with germ cells from closely related donor fish (either endangered or more commercially valuable species). One example in Japan involves using edited grass puffer fish (Takifugu alboplumbeus) to serve as surrogate broodstock for the more valuable tiger puffer (T. rubripes).
Enhancing resistance to various pathogens may seem like fertile ground for the application of CRISPR in aquaculture, but disease resistance is a complex phenomenon involving the interaction of host species with both pathogens and the production environment, and many genes are usually involved. One research focus that shows commercial promise involves identifying genes that confer enhanced sea lice resistance in Pacific salmon species and then establishing similar attributes in Atlantic salmon through editing.
Japanese researchers reported in 2018 on the use of CRISPR techniques for the development of a line of red sea bream with enhanced muscle mass and a relatively short overall body length. Mutations were created by deletions in the Pm-mstn myostatin gene, and no exogenous genetic constructs were used. Compared to non-edited bream, the line exhibited a 16 percent increase in edible muscle tissue, and within two years a stable breeding population was established. As is the case in many other efforts with aquaculture species, this work built on prior research in laboratory populations of medaka and zebrafish. Two years later, Chinese investigators reported on a strain of yellow catfish (Pelteobagrus fulvidraco) that was also produced through editing of the myostatin gene. At 210 days post-fertilisation, fish from the edited line were 37 percent heavier than regular P. fulvidraco, with increased muscle mass.
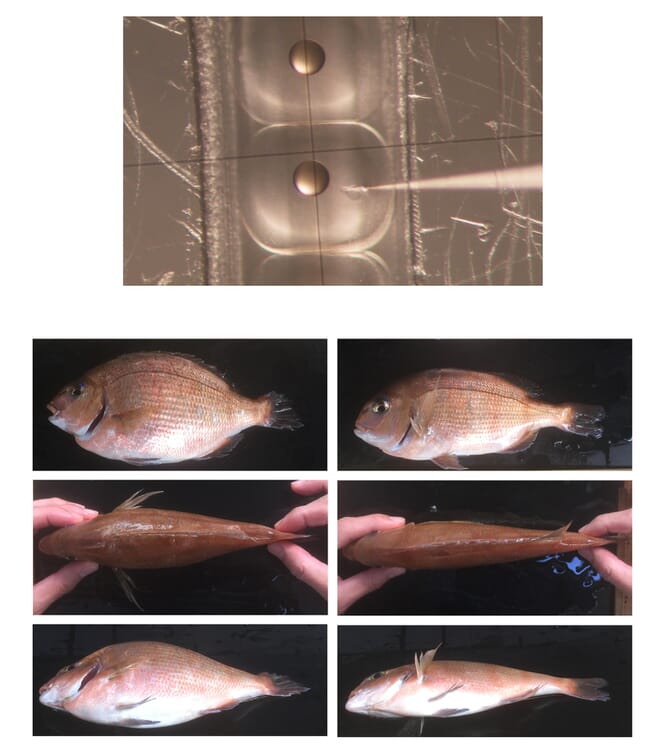
© Dr Masato Kinoshita, Kyoto-University and Dr Keitaro Kato, Kindai University
Editing myostatin genes has also been reported to significantly increase muscle mass in common carp, olive flounder, blunt snout bream, Sea bream, mud loach and channel catfish over the past five years, and scientists are now beginning to look at other genes in efforts to enhance growth rates in aquatic species. In the past several months, another group of researchers from China reported on the use of CRISPR to knock out the PI3K gene in Gibel carp. Disruption of this gene improves insulin sensitivity in mammals, but edited carp exhibited no alteration of plasma and hepatic glucose levels or uptake. They did, however, have improved somatic growth and feed conversion efficiency. Another recent study in China established that deletion of the t1r1 gene significantly improved acceptance of plant proteins in zebrafish.
Colouration and colour patterns are also important considerations in some aquaculture species. In China, researchers used CRISPR to disrupt carotenoid transport genes to alter red and white colour patterns in ornamental common carp. Another group used the technology to alter two agouti signalling protein (ASIP) genes to eliminate black patches in Oujiang common carp. Also in China, in a study to be published shortly, researchers used CRISPR to knock out the tyrosinase gene in a line of red tilapia. Once a true breeding population was established, continuous production of uniformly red fish with no black pigment was possible. And Israeli scientists recently published results (in The CRIPSR Journal) detailing the use of CRISPR to produce true albino Nile tilapia (pink eyes and all). They disrupted the slc45a2 gene, which mediates melanin biosynthesis, to produce zygotes with up to 99 percent albinism, including lack of melanin in the eyes.
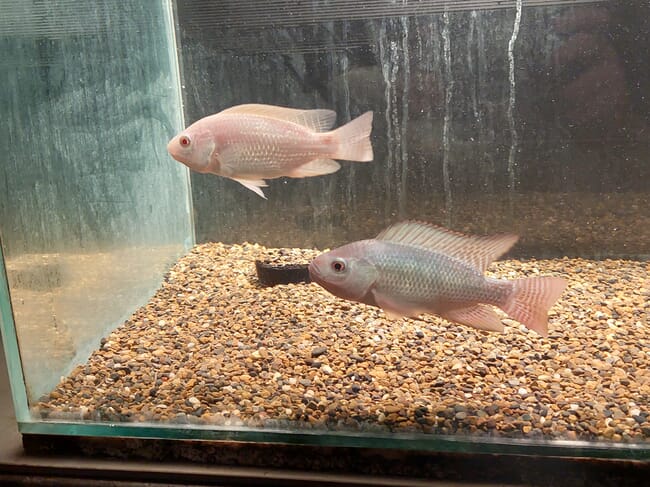

© Dr Jakob Biran
Use of CRISPR technology has also been demonstrated in crustaceans. Researchers from China reported in 2016 on the first genome editing of a decapod, the ridgetail white prawn Exopalaemon carinicauda. The primary objective of the work was to clarify the function of the chitinase compound EcChi4, by knocking out the gene responsible for its production. Since this species carries the fertilised eggs prior to hatching, one-cell stage embryos were collected from newly spawned females, stored in sterilised seawater at 4 °C to halt further development and then microinjected. After 15 days of artificial incubation at room temperature, the shrimp hatched and were raised through the juvenile phase. Genomic DNA was extracted from both mysis larvae and juveniles to determine whether CRISPR methods resulted in mutation of the target gene. Results indicated different mutations were induced in the target gene, and that mosaic shrimp had been produced. The rate of mutation was calculated at 7.3 percent, and when individuals that were heterozygous for the induced mutation were crossed with normal shrimp, Mendelian inheritance (with a simple pattern where genes segregate into gametes at equal frequencies) was observed. When heterozygous offspring of the mutant shrimp were crossed, roughly one quarter of their offspring were homozygous for the mutation. In the case of EcChi4 in E. carinicauda, the induced mutation was heritable and did not influence survival or growth. Other crustaceans that have successfully been modified using CRISPR include Daphnia magna and the amphipod Parhyale hawaiensis.
To approach a CRISPR-based strategy, some knowledge of an organism’s genome is required. While few and far between, some bivalve genomes are tentatively available for this type of research, including those of the Pacific oyster, eastern oyster, pearl oyster, Sydney rock oyster, Mediterranean mussel, Philippine horse mussel, king scallop and Yesso scallop. Nonetheless, work on bivalve gene editing has been quite difficult to date. Researchers have reported on the use of CRISPR targeting the myostatin gene in the Pacific oyster, with some success using microinjection methods.
Looking ahead
For the time being, applying CRISPR to aquatic species will continue to be a pursuit of academic and research institutions. It’s one thing for a university to develop the molecular technology and expertise required, but for private concerns to establish such high-tech capabilities a tremendous investment in equipment and staff will be required. Nevertheless, CRISPR is cheaper and more precise than other gene editing alternatives and usually provides better results.
Future applications and implications for CRISPR-related research in aquaculture species are currently debated, sometimes passionately, in many scientific and social contexts. One thing we can be certain of is the steady progress in our understanding of the genomes and complex physiological interactions of many important aquatic species, which in turn will allow more precisely targeted gene editing to improve production characteristics. And all with the potential to actually minimise genetic impacts on wild fisheries.







