
A project that uses genetic markers has identified family differences in farmed Pacific oyster for resistance against herpes virus. Two genetic markers explained about half of the genetic variation for survival against the virus. The knowledge can help industry to select oysters efficiently and effectively for improving survival.
Background
The Pacific oyster (Crassostrea gigas) is a highly important shellfish species, accounting for roughly 98 percent of global oyster aquaculture. Infectious diseases pose a significant threat to sustainable production, with high economic losses due to mortalities.
The Ostreid herpesvirus type 1 (OsHV-1) is highly contagious, with a relatively short lifecycle and can cause up to 100 percent mortalities in infected oysters. Juvenile oysters (2–5 cm/5–20 g) are much more susceptible than adult oysters (>5 cm/>20 g). An outbreak is characterised by a reduction in feeding and swimming activities, along with sudden high mortalities. OsHV-1 virus can infect several other bivalve species – including blue mussel, European flat oyster, wedge clam and king scallop – demonstrating the robustness and adoptability of OsHV-1 in wide range of hosts, and making it highly threatening once detected in a system.
Genes affecting survival
The analyses, by scientists from Nofima and Ifremer, found family differences for resistance against OsHV-1 virus. Hence, some families showed better survival than others and the selection of individuals (as parents of next generation) from resistant families using genetic markers would efficiently improve survival against an OsHV-1 outbreak.
The results produced in this study are a part of EU-funded project VIVALDI (H2020 program, n°678589). The oyster population was produced by specific F2 cross, originating from the breeding nucleus of Ifremer, in France, by the admixing of resistant and susceptible grandparents. The study utilised a sampling opportunity following a natural field outbreak of OsHV-1 in the sea.
Approach
At the end of the OsHV-1 outbreak, the collected tissue samples of the dead and the surviving individuals were selected in equal numbers, in a bid to increase the power to detect genetic markers affecting the survival against the virus. The selected shellfish were genotyped using a 40,000 single-nucleotide polymorphism (SNP) array (Thermo-Fisher, AXIOM). The data (survival phenotype and genotype) were analysed to detect SNP markers and/or quantitative trait loci (QTLs) associated with the survival against OsHV-1, which led to the detection of two significant genomic regions.
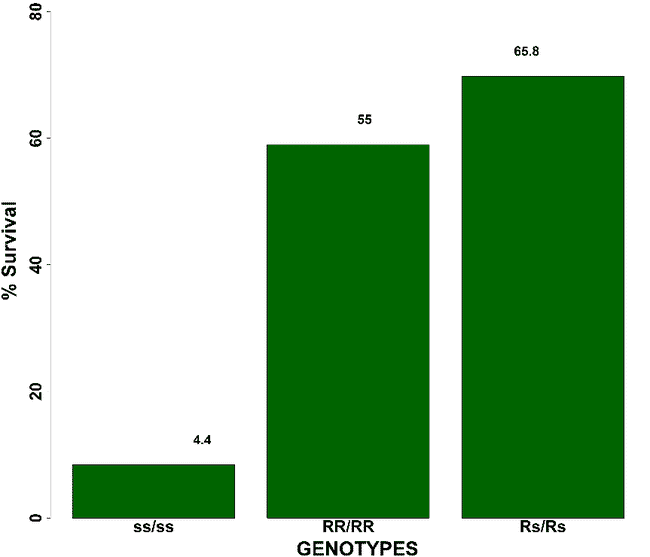
Percentage survival against OsHV-1 virus across three different QTL genotypes. The survival values are calculated within each QTL genotypes. The allele “s” at the QTL position represents unfavourable mutation, which makes the animal susceptible while the mutation “R” is favorable which makes the individual resistant. The QTL genotypes “Rs/Rs” seems to promise the highest survival during OsHV-1 outbreak due to phenomenon of over-dominance.
Impact
The two markers (top marker from each region) explained about 50 percent of the genetic variation for survival against OsHV-1, and therefore can help industry to select oysters efficiently and effectively for improving survival against OsHV-1. The impact of the detected markers is similar to markers identified against cardio (CMS) and the pancreatic (PD) diseases in Atlantic salmon, which are now used/adopted as markers/QTL products in the salmon farming industry. Therefore the QTL markers identified for survival against OsHV-1 have a strong potential for the Pacific oyster industry.
Jean-Baptiste Lamy, a scientist from Ifremer, is very pleased with the findings. He thinks that the results of this study on genetic variation, and the QTL-findings which explain large proportions of the genetic variance, will convince oyster breeding companies to implement more carefully designed breeding programmes, and test advanced – but relatively capital intensive and efficient – solutions, such as marker assisted and/or genomic selection. In addition, the results from this study should inspire more studies on other complex diseases caused by Vibrio aesturianus, and the simultaneous infections between viruses and bacteria, that threaten global oyster production.







