These studies have yielded almost 500 streptococcal isolates recovered from tilapia at approximately 50 sites in 13 countries over the last 8 years. The isolates were identified using standard biochemical and bacteriological identification methods and subsequently analyzed by cluster analysis (unweighted pair-group average based on percent disagreement). Interestingly, of the nearly 500 streptococcal isolates recovered from tilapia, 82 per cent were identified as Streptococcus agalactiae and 18 per cent were identified as Streptococcus iniae.
S. iniae is a significant fish pathogen causing disease and mortality in many marine and freshwater cultured fish species in tropical and sub-tropical areas. Vaccines to combat S. iniae infection in a variety of fish species including tilapia are available, and there is a large body of literature on the pathogenesis of this organism in a variety of fish species.
Considerably less information is available for fish-pathogenic S. agalactiae. Although more commonly associated with disease in human and bovine hosts, fish-pathogenic S. agalactiae has been documented from as early as 1966, when a non-hemolytic Group B Streptococcus was identified as the cause of two epizootics in golden shiners (Notemigonus crysoleucas). Today, with the intensification of aquaculture, we find S. agalactiae to be a significant cause of mortality and morbidity in both marine and freshwater cultured species and particularly in tilapia.
| Table 1. Major streptococcal pathogens of tilapia and their prevalence in the Asia-Pacific region | |||
|---|---|---|---|
|
Global prevalence
(as per cent of total streptococcal isolations)* |
|||
|
S. agalactiae Biotype 1
|
26
|
||
|
S. agalactiae Biotype 2
|
56
|
||
|
S. iniae
|
18
|
||
|
* Data generated by Intervet/Schering-Plough Animal Health, Singapore
|
|||
Detailed analysis of our tilapia S. agalactiae isolates suggests the presence of two distinct clusters, which differ in a variety of biochemical and phenotypic characteristics. We refer to these distinct clusters as biotypes; on this basis, we differentiate between typically beta-hemolytic “classical” S. agalactiae (hereafter referred to as S. agalactiae Biotype 1) and typically non-beta-hemolytic S. agalactiae (hereafter referred to as S. agalactiae Biotype 2). These latter strains were previously classified as S. difficile /S. difficilis but have subsequently been reclassified as non-hemolytic variants of S. agalactiae.
In our epidemiological surveys to date, 26 per cent of all streptococcal isolates of tilapia were found to be S. agalactiae Biotype 1 and 56 per cent were identified as S. agalactiae Biotype 2 (Table 1). The significance of this observation and how it relates to the development of S. agalactiae vaccines for tilapia will be explored in this paper.
Materials and methods
Bacterial isolates recovered from diseased tilapia were identified as S. iniae or S. agalactiae Biotype 1 or Biotype 2 using a variety of standard biochemical and phenotypic tests. The test results were analyzed by cluster analysis (unweighted pair-group average based on percent disagreement). Alternatively, identification was made using species- and/or biotypespecific PCR.
Experimental S. agalactiae Biotype 1 or Biotype 2 vaccines were manufactured as water-in-oil emulsions containing inactivated biotype-specific bacterial antigens and metabolizable non-mineral oil. The vaccines were given as a single dose to fish weighing 15 g or more.
The efficacy of the biotype-specific vaccines was evaluated in laboratory trials following intraperitoneal vaccination of 15 g tilapia. Age-matched, unvaccinated fish from the same cohort and origin were maintained as controls.
At various time points after vaccination, vaccinated and control fish were challenged by intraperitoneal injection with virulent heterologous S. agalactiae Biotype 1 or Biotype 2 strains. Fish were observed for 14 or 15 days post-challenge and mortality was recorded daily. Post-mortem recovery of the challenge organism was performed on fish that died during the observation period and on all surviving fish at the end of the observation period.
Results
To date, we have identified the species and, for S. agalactiae, the biotype of almost 500 streptococcal isolates gathered from approximately 50 sites in 13 countries. This is not a true epidemiological analysis since some sites were visited on multiple occasions and are therefore overrepresented in the data set, but this nonetheless represents a detailed analysis of the impact of streptococcal disease in tilapia.
As mentioned above, 26 per cent of all streptococcal isolates of tilapia were found to be S. agalactiae Biotype 1 and 56 per cent were identified as S. agalactiae Biotype 2; 18 per cent were identified as S. iniae.
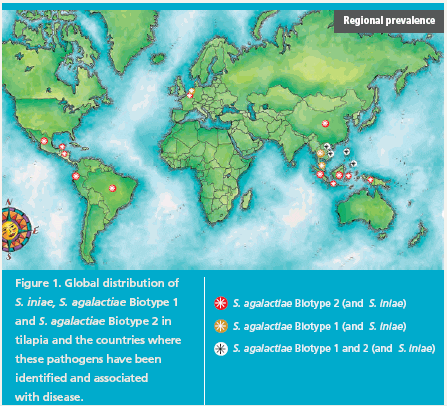
It is intriguing that our analysis further suggests that the S. agalactiae biotypes are present in distinct geographical zones (Figure 1).
From our sampling of tilapia worldwide, S. agalactiae Biotype 2 is the most prevalent and geographically diverse of the streptococcal pathogens. In Asia, we find S. agalactiae Biotype 2 in China, Indonesia, Vietnam and the Philippines, and in Latin America, we have found it in Ecuador, Honduras, Mexico and, most recently, in samples from Brazil.
In contrast, we find S. agalactiae Biotype 1 to be the dominant streptococcal pathogen of tilapia in Thailand, Malaysia and Singapore. S. iniae is often found in association with S. agalactiae Biotypes 1 or 2 in China, Ecuador, Honduras, Indonesia, the Philippines and Thailand. Only in the Philippines and Vietnam have we observed S. agalactiae Biotype 1, S. agalactiae Biotype 2 and S. iniae in tilapia in the same country.
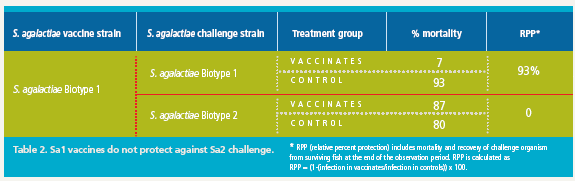
It is known that S. iniae vaccines do not provide protection against S. agalactiae infection. To determine if our classification of the fish pathogenic S. agalactiae has consequences for the development of vaccines to control this devastating disease, we assessed in a laboratory challenge the ability of biotype-specific vaccines to protect against lethal challenge with S. agalactiae Biotype 1 or Biotype 2 strains. The results of representative experiments are shown in Figures 2 and 3 and are summarized in Tables 2 and 3.
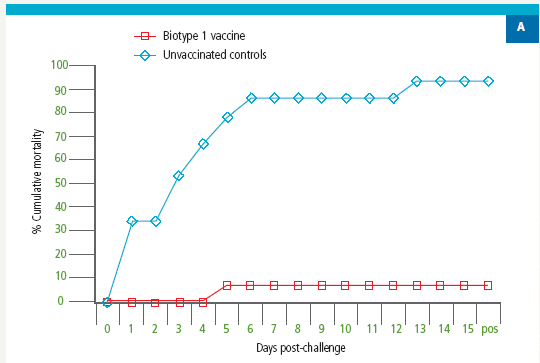
Tilapia vaccinated with an experimental S. agalactiae Biotype 1 vaccine were protected against lethal challenge with a virulent S. agalactiae Biotype 1 strain (Figure 2A). However, no protection against challenge with a virulent Biotype 2 strain was observed in the Biotype 1-vaccinated fish (Figure 2B).
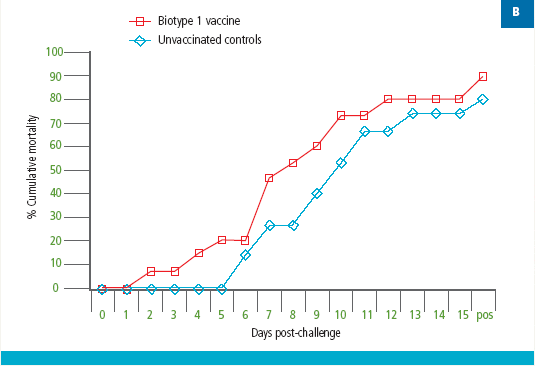
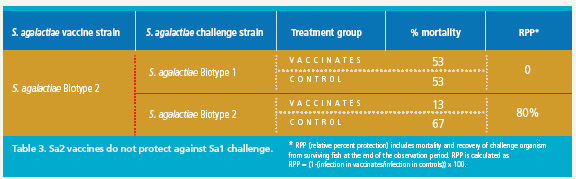
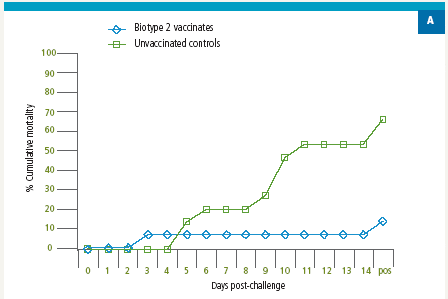
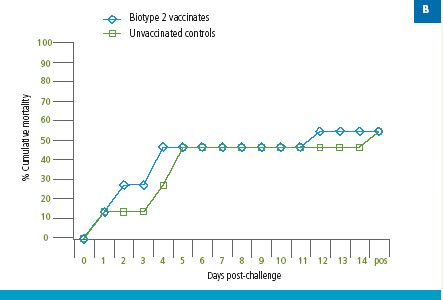
Similarly, fish vaccinated with a S. agalactiae Biotype 2 vaccine were protected against lethal S. agalactiae Biotype 2 challenge (Figure 3A); however, there was no protection against challenge with a virulent Biotype 1 strain in the Biotype 2-vaccinated fish (Figure 3B). Thus, vaccination with biotype-specific bacterin vaccines induces biotype-specific protection against mortality caused by S. agalactiae.
Conclusions
Our data suggest that S. agalactiae, and to a lesser extent S. iniae, are the principal agents of streptococcosis in tilapia. Detailed analysis of our tilapia S. agalactiae isolates suggests the presence of two biotypes, which differ in a variety of biochemical and phenotypic characteristics.
In our experience, these two S. agalactiae biotypes cause subtly distinct disease syndromes. S. agalactiae Biotype 1 infects fish throughout the production cycle from juvenile to grow-out, while S. agalactiae Biotype 2 causes disease predominantly in larger fish.
Moreover, and most significantly from a health-management perspective, we have demonstrated that immunity is biotype-specific.
To our knowledge, there are no obvious geographical, physiological or environmental explanations for the country-to-country distribution of S. agalactiae Biotypes 1 and 2. It would be prudent, however, to consider that the distribution might change in time, probably through trade in live fish.
December 2009

