Various medicinal treatments have been devised to control sea lice numbers (Pike & Wadsworth 2000), but the risk of resistance developing to these treatments has led to increasing interest in the use of cleaner fish such as wrasse, which can live within salmon cages and pick sea lice from the skin of salmon (Bjordal 1991).
Although adopted widely in the 1990s (Treasurer 1996), the possible problems of wrasse harbouring bacterial pathogens (Treasurer 2012) and concerns over the use of large numbers of wild-caught wrasse (Treasurer 1996) led to a decline in their use in favour of easily administered and effective in-feed treatments. However, there is renewed interest in the use of wrasse (www.eco-fish.org) due to developing resistance against various medicines (reviewed by Torrissen et al. 2013), and rearing of several species of wrasse is in progress in Norway and Scotland. Here, we report on the digestive tract bacterial flora of ballan wrasse, Labrus bergylta Ascanius, and goldsinny wrasse, Ctenolabrus rupestris (L.), reared at the Ardtoe Marine Laboratory, Scotland. Because vibrios dominated the flora of the digestive tract of marine fish larvae in previous studies (Reid et al. 2009) and vibrios have been reported as pathogens of wrasse (Gravningen, Kvenseth & Hovlid 1996; Bergh & Samuelsen 2007), this study examines the vibrio species found in the gut of larval wrasse and assesses the potential impact on culture of labrids. Samples were taken from one batch of ballan wrasse larvae and two batches of goldsinny wrasse larvae under commercial rearing conditions.
Wrasse larvae were reared using protocols given in Eco-fish (2012). Briefly, larvae were initially reared with microalgal addition and offered live feed of enriched rotifers for the first 28 days post-hatch (dph) followed by transition to Artemia. In the first sampling, six ballan wrasse larvae (150 dph; mean length, 35.1 mm; SEM = 0.87 mm) were removed from one 1300-L volume rearing tank (temperature, 10–12 °C and salinity, 32–34 per cent) in which mortalities of approximately 2 per cent per day were occurring. At this time, larvae received Artemia feed at 12-h intervals, with concentrations maintained >0.5 Artemia mL-1. For the second sampling, five goldsinny wrasse larvae (102 dph; mean length, 15 mm) were used. Temperature, salinity, rearing protocols and daily mortalities were as above. For the third sampling, five goldsinny wrasse (150 dph; mean length, 39.2 mm) were used. These were from the same tank as the wrasse sampled on day 102, and larvae were fed Artemia daily supplemented with Nofima dry wrasse diet (200–300 lm diameter). The water temperature range was 10.5–12.5 °C, salinity was 32–34 per cent, and mortalities were approximately 1.5 per cent per day.
Bacteriological sampling from larvae was as described by Munro, Barbour & Birkbeck (1993). Briefly, larvae were terminally anaesthetized by immersion in 0.1 per cent (w/v) methacaine tri-sulphate (MS222; Alpharma) solution in autoclaved sterile sea water (SSW) and then surface sterilized in 0.1 per cent (v/v) benzalkonium chloride in SSW for 30 s (Munro et al. 1993).
The fish were then rinsed in two changes of SSW, and soft tissues from two fish were dissected from the head and notochord, combined and homogenized in 1.5-mL microfuge tubes with micropestles (Eppendorf). The homogenate was made up to 1 mL volume with SSW and 10-fold serially diluted in SSW before appropriate dilutions were spread onto duplicate tryptone soy agar (TSA, Oxoid) + 2 per cent NaCl plates that were incubated at 20 °C for up to 7 days. After initial colony-forming unit (cfu) counts, all colonies from the 10-2 dilution plates were inoculated into 100 μL volumes of marine broth (Difco) in the wells of a 96-well tissue culture plate.
Broth cultures were incubated at 20 °C overnight, before addition of 100 lL 40 per cent glycerol in 3 per cent sea-salts solution to each well for storage at 80 °C. Goldsinny wrasse larvae at 102 dph were sufficiently small for complete larvae to be homogenized but larvae from 150 dph were processed as mentioned above. Water from the ballan wrasse rearing tank was also analysed as described by Verner-Jeffreys et al. (2003). Bacteria from the master plates stored at 80 °C were replica plated onto selective agar media using a replica plater (Reid et al. 2009).
After culture on TSA + 2 per cent NaCl, colonies were tested for bioluminescence, swarming, oxidase, catalase, haemolysis on TSA + 2 per cent NaCl agar + 5 per cent sheep blood, growth on thiosulphate citrate bile salts (TCBS) agar, utilization of sucrose on TCBS agar, growth at 4, 20 and 35 °C, β-galactosidase, aesculin hydrolysis, production of acid from arabinose, glycerol, maltose, mannose, mannitol and arbutin (Furniss, Lee & Donovan 1978; Cowan et al. 1993). Data from the above tests were recorded in binary fashion and processed by the least squares method using the program CLUSTAN Graphics 4.0 (Clustan) that was run on a PC.
PCR and determination of the 16S rRNA gene partial sequence of representative isolates were performed as described by Reid et al. (2009) followed by BLAST searches (Altschul et al. 1997) of the GenBank database.
The reactions of the type strains of V. ichthyoenteri (NCIMB 13441) and V. scophthalmi (NCIMB 13623) and four putative isolates of V. ichthyoenteri were compared for arginine dihydrolase, lysine decarboxylase and ornithine decarboxylase activities (Furniss et al. 1978) and in Biolog GN plates (TechnoPath Distribution Ltd) as described by Verner- Jeffreys et al. (2003).
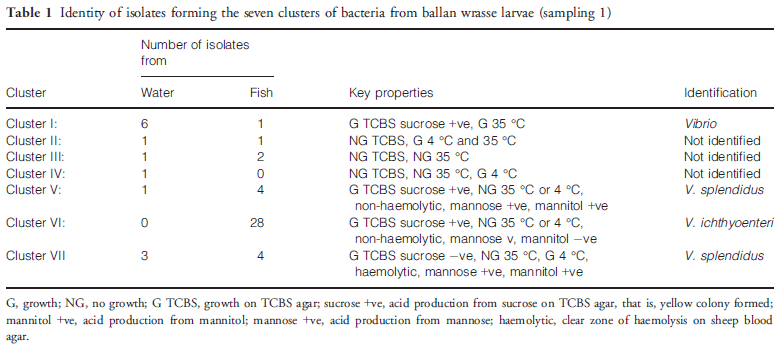
Ballan wrasse larvae contained 3.2 9 104 cfu culturable bacteria per larva and the tank water contained 320 cfu mL-1. Some slow-growing colonies did not survive replica plating, and in total, 53 isolates were analysed (13 from water and 40 from larvae). Of these, 47 grew on TCBS agar, a medium that is largely selective for Vibrio, and these bacteria were considered presumptively as members of the Vibrionaceae. Cluster analysis divided the isolates into seven groups (Table 1), 4 of which were presumptive vibrios. Clusters I–IV were not analysed further as they were mainly from water. Five isolates from each of phenons V and VII were tested for agglutination using an antiserum to V. splendidus DMC-1 (Macpherson 2004) and all were agglutinated, whereas isolates from phenon VI were not agglutinated. Partial sequencing of the 16S rRNA gene of a total of five isolates from phenons V to VII led to confirmation of phenons V and VII as V. splendidus (99.8 per cent identity over 550 nt) and phenon VI as V. ichthyoenteri (99.7 per cent identity over 650 nt).
A more extensive biochemical comparison was made of putative V. ichthyoenteri isolates with the type strains of V. ichthyoenteri and V. scophthalmi. No firm conclusions were drawn from BIOLOG GN analysis as V. ichthyoenteri NCIMB 13441T, V. scophthalmi NCIMB 13623T and the 4 wrasse isolates tested reacted with only 7, 14 and 13 of the 95 substrates, respectively. However, in four key differentiating characteristics between V. ichthyoenteri and V. scophthalmi, the 4 wrasse isolates reacted exactly as for V. ichthyoenteri (growth in 6 per cent NaCl, utilization of maltose, lack of growth at 35 °C, lack of arginine decarboxylase activity) with V. scophthalmi giving the opposite reactions.
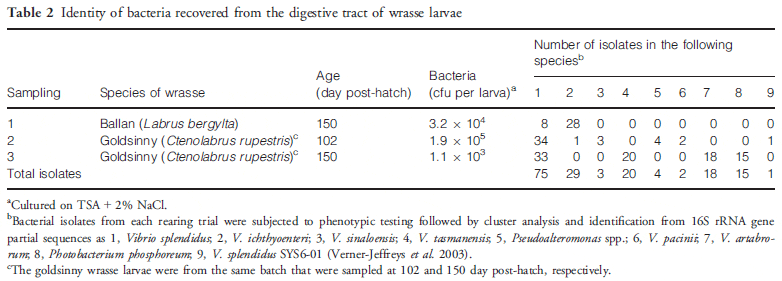
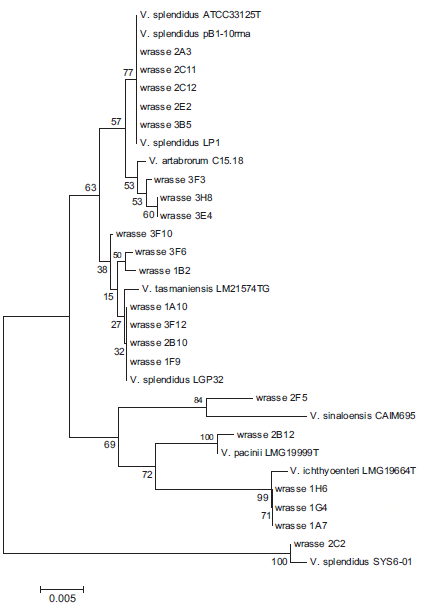
The results of bacteriological sampling of goldsinny wrasse larvae are summarized in Table 2 where they are compared with results for ballan wrasse rearing. The goldsinny wrasse larvae contained 1.9 X 105 and 1.1 X 103 cfu per larva on days 102 and 150, respectively, culturable on TSA + 2 per cent NaCl agar plates. In the sample from day 102 (experiment 2), 45 colonies from one of the 10-3 dilution plates were successfully subcultured into TSB + 2 per cent NaCl in a 96-well plate. Of these, 41 grew on TCBS agar and were presumed to be Vibrionaceae. In the sample from 150 days (experiment 3), all 96 isolates cultured from these larvae grew on TCBS agar and were presumptive Vibrionaceae. Following cluster analysis and BLAST searches of the 16S rRNA gene sequences of representative isolates, a neighbourjoining phylogenetic tree was produced for the isolates and reference strains of the closest matches identified from BLAST (Fig. 1). From this analysis, the 117 isolates from all three sets of samples were dominated by V. splendidus (75 isolates) that was found in all three samples with V. ichthyoenteri (29 isolates) found in two samples. Other vibrios and Pseudoalteromonas were only found in one sample. The bacterial load of larval feed (Artemia and dry feed) was also analysed, but as only two colonies of a total of 128 from Artemia grew on TCBS agar (non-sucrose fermenting) and none of 21 from dry feed grew on TCBS agar, the colonies were not analysed further.
The samples taken were intended to identify the principal bacteria present under normal rearing conditions and the presence of any potential pathogens (Treasurer 2012). Vibrio splendidus and closely related bacteria dominated the intestinal bacterial flora of goldsinny wrasse larvae, and this microorganism is a pathogen of wrasse (Jensen et al. 2003; Bergh & Samuelsen 2007) and turbot (Gatesoupe, Lambert & Nicolas 1999; Thomson et al. 2005) causing damage to the intestinal tract of larvae by producing an aerolysin-like enterotoxin (Macpherson, Bergh & Birkbeck 2012).
Phylogenetic analysis (Fig. 1) revealed that most sequenced isolates clustered with V. splendidus and the closely related organisms V. tasmaniensis and V. artabrorum. Similar results were obtained with a nearest-neighbour phylogenetic tree (results not shown). Bacteria identified as V. splendidus vary considerably in biochemical properties and 16S rRNA gene sequence. Thompson et al. (2005) estimated that a natural population of bacterioplankton contained over 1000 different genotypes of V. splendidus, and differences in 16S rRNA sequence between multiple copies of the gene within one vibrio can be greater than between closely related species (Michon et al. 2010). Apart from these differences within V. splendidus, closely related species such as V. artabrorum (Dieguez et al. 2011) and V. tasmaniensis (Thompson, Thompson & Swings 2003) have been described within the ‘splendidus clade’.
Two other potential pathogens were detected in this study. Vibrio ichthyoenteri was originally isolated as a pathogen of Japanese flounder larvae (Ishimaru, Akagawa-Matsushita & Muroga 1996) and V. pacinii is noted as a pathogen of salmon (Gomez-Gil et al. 2003). Both these species were also found in cod larvae (Reid et al. 2009) at this hatchery. In Fig. 1, the isolates identified as V. ichthyoenteri, V. pacinii and V. sinaloensis were clearly separated from the broad ‘clade’ of V. splendidus isolates.
In summary, the majority of the bacteria identified from the intestinal tract of wrasse larvae were V. splendidus or closely associated bacteria, and given that many V. splendidus strains are pathogenic for marine fish larvae, it is likely that some isolates of V. splendidus, V. ichthyoenteri and V. pacinii could be pathogenic if the general fitness of larvae is depressed. Routine monitoring of the intestinal tract bacterial flora of wrasse larvae is recommended to survey the population of vibrios and haemolytic bacteria and to identify health risks to larvae.
May 2014



