Control of disease is complex and relies heavily on a combination of pathogen detection, disease diagnosis, treatment, prevention and general health management. Rapid disease diagnosis and vaccination play a crucial part in this and the team from the Aquatic Vaccine Unit, Institute of Aquaculture, led by Professor Sandra Adams have made a significant contribution to both of these areas over the last twenty years.
Research in the Aquatic Vaccine Unit focuses on fish disease control through the development of novel rapid diagnostic tests (both molecular and antibody based) and vaccines for both fresh water and marine cultured fish species. In addition, basic research focuses on the elucidation of host pathogen interactions and the immune system of fish. Numerous vaccines, antibodies and kits have been/are being commercialised from research projects performed at Stirling. The Aquatic Vaccine Unit is also an active partner in the Scottish Fish Immunology Centre with partners in Aberdeen (Aberdeen University and Marine Scotland immunology laboratories), promoting the training of immunologists. Moreover, it has close links with local schools, taking Nuffield Bursary placements every year.
Fish Vaccines
Vaccination has made a huge impact in reducing the risk of furunculosis in salmon in Scotland and Norway. This is turn led to a reduction in the use of antibiotics that has been sustained, and an increase in productivity as a result of vaccination. Vaccines developed in the Aquatic Vaccine Unit are part of this success with a furunculosis vaccine being developed in a joint programme between Marine Scotland in Aberdeen and Stirling. Other vaccines developed at Stirling in the late 1980’s and 1990’s include enteric redmouth (ERM), vibriosis and pasteurella vaccines, with many others in the pipeline over recent years.
Vaccination has been extremely successful in reducing the disease risk in fish, however, biological, scientific and technical restrictions still prevent the production of commercial vaccines for most economically significant fish diseases. Knowledge of the sequence of pathogen genomes, gene function and derived products now allows innovative approaches for vaccine development using genetic, immunology and chemistry/ physiology-based technologies.
These are the kind of approaches now being taken by Stirling for vaccine development. In addition, novel vaccine antigen identification methods are being devised using combinations of various techniques such as genomics, proteomics, knockout technologies and epitope mapping. In other words fish vaccine development has become much more sophisticated in recent years. These novel vaccines are being developed because the simpler approach of using inactivated whole cell vaccines does not always suceed, and attempts at attenuated vaccines in general have not been encouraged from a safety point of view.
In order to develop an effective vaccine the protective antigens need to be identified and their protective response confirmed in the host species. Research in the Aquatic Vaccine Unit has been focusing on improving methods to identify which serotypes to include in traditional whole cell vaccines, as well as developing new methods to identify potential vaccine antigens. Four examples are presented here where vaccine antigens have been sucessfully identified at Stirling using different technologies: rainbow trout fry syndrome (RTFS), vibriosis in cod, Aeromonas hydrophila in carp and betanodavirus in sea bass. Such methods can be used for the development of many other vaccines.
Many of the current fish bacterial vaccines are whole cell vaccines where the isolates included are simply collected from disease outbreaks and are incorporated on that basis.
Inclusion of all serotypic variants is important but in many cases serotyping systems are not robust. Thus, a new serotyping system was developed for Flavobacterium psychrophilum so that representative isolates could be selected from each serotype. The F. psychrophilum vaccine developed by Dr Alison Morgan, Kim Thompson and Sandra Adams is currently on field trial with a view to commercialisation.
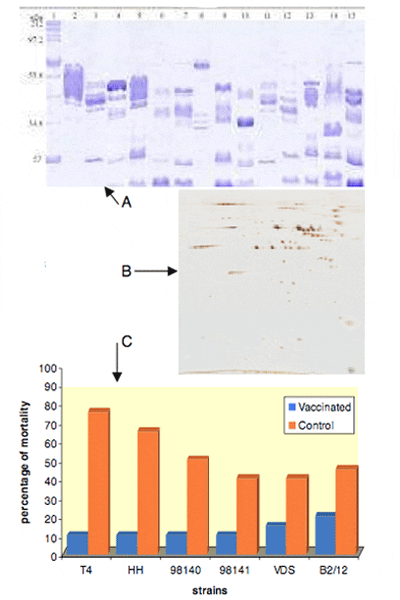
A. Antigenic diversity of outer membrane profiles of hydrophila isolates by SDS polyacrylamide gel electrophoresis (PAGE)
B. Western blotting sera from infected carp on to 2D SDS gels identified potential vaccine antigens
C. Vaccination with the recombinant vaccine protects against a variety of field isolates of A. hydrophila to different extents
Farmers have found it difficult to treat A. hydrophila infections in fish due to the resistance of this pathogen to a number of antibiotics. Thus, researchers have been trying to develop a vaccine but this has proven difficult, mainly because of the differences in the antigenic (bacterial components) expression between strains of A. hydrophila. A new approach using immunoproteomics was used to develop a recombinant vaccine that would protect against a wide range of A. hydrophila strains.
Salmon vaccines from the past are based on inactivated (whole cell) cultures of the pathogenic organism grown in vitro. Some of the vaccines gave good protection, however, many pathogens have been shown to ‘switch off’ important protective antigens when cultured in vitro and so many of these vaccines failed. In such cases alternative methods of culture (e.g. the inclusion of iron chelating agents) were required so that expression of the important ‘protective’ antigens were induced, e.g. in the development of a furunculosis vaccine.
This was accomplished by Saravananne Poobalane, Kim Thompson and Sandra Adams who implanted live A. hydrophila into fish (enclosed in sealed chambers), i.e. IVET. Application of sera from fish (that had been infected with A. hydrophila and then recovered) in Western blot analysis on one and two-dimensional gel electrophoresis (immunoproteomics) pinpointed potential vaccine candidates and a recombinant vaccine against A hydrophila was produced and shown to be protective in carp (Cyprinus carpio) (Fig. 2). This vaccine will be taken forward to commercialisation. Recombinant and DNA vaccine technologies are powerful tools for vaccine development as these enable the isolation of potential protective antigens from suppressive ones.
Epitope mapping is a useful technology for the identification of protective antigens, particularly for viruses. The Pepscan procedure was used by Janina Costa, James Bron, Kim Thomspon, William Starkey, Randolph Richards and Sandra Adams to identify betanodavirus B-cell epitopes recognized by neutralizing mouse monoclonal antibodies (MAbs) and serum samples obtained from European sea bass, Dicentrarchus labrax, naturally infected with betanodavirus. Pepscan was performed with a panel of thirty-four 12-mer synthetic peptides that mimicked the entire betanodavirus capsid protein. Sea bass serum samples reacted strongly with three regions of the capsid protein and one of these regions was also recognized by neutralizing MAbs and therefore could potentially be used towards the development of nodavirus vaccines.
Rapid, accurate diagnosis of disease is essential for effective outbreak control. Once the causative agent has been identified appropriate advice can be given to neighbouring farmers/ trading partners. Prompt action in the early stages of any disease problem can have an enormous impact on the scale of the outbreaks. Rapid diagnostic methods therefore provide powerful tools during emergency management.
A variety of methods are available to detect pathogens in both fish and in the aquatic environment. These include traditional diagnostic methods and a diverse range of immunological and molecular methods. A variety of novel rapid diagnostic methods (immuno and molecular diagnostics) have been developed for use in aquaculture over the last 20 years in the Aquatic Vaccine Unit. The availability of standardised reagents (e.g. antibodies for immunodiagnostics) is crucial if comparable tests are to be run in different laboratories.
The Aquatic Vaccine Unit was instrumental in developing a wide range of monoclonal antibodies (mAbs) for this purpose and a spin-out company, Aquatic Diagnostics Ltd (www.aquaticdiagnostics.com), was set up in 2001 by Kim Thompson and Sandra Adams following a SMART Biotechnology award to market these reagents worldwide.

